Model Evaluation¶
Evaluate fitted models against ground truth: convergence diagnostics, mutation effect accuracy, shift sparsity, replicate correlations, and variant phenotype predictions.
Outline
Convergence diagnostics — check converged status, plot loss trajectories
Mutation effect accuracy — beta and shift vs ground truth, save
model_vs_truth_beta_shift.csvShift sparsity — fraction of shifts exactly zero, save
fit_sparsity.csvReplicate correlations — lib_1 vs lib_2 mutation parameters, save
library_replicate_correlation.csvVariant phenotype predictions — predicted vs true variant phenotypes, save
model_vs_truth_variant_phenotype.csvAdditional visualizations — latent vs functional score, pairplot, violin by variant class, phenotype vs lambda
[1]:
import warnings
warnings.filterwarnings("ignore")
import itertools
import os
import pickle
import sys
from collections import defaultdict
sys.path.insert(0, "notebooks")
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import seaborn as sns
import multidms
import multidms.plot
from multidms.model_collection import ModelCollection
from _common import (
load_config,
reconstruct_simulators,
)
%matplotlib inline
[2]:
config_path = "config/config.yaml"
[3]:
# Parameters
config_path = "config/config.yaml"
[4]:
config = load_config(config_path)
sim = config["simulation"]
fit_config = sim["fitting"]
seed = config["seed"]
train_frac = config["train_frac"]
output_dir = sim["output_dir"]
lasso_choice = sim["lasso_choice"]
os.makedirs(output_dir, exist_ok=True)
print(f"Output directory: {output_dir}")
Output directory: results
Load data¶
[5]:
fit_collection_df = pickle.load(
open(os.path.join(output_dir, "fit_collection.pkl"), "rb")
)
mut_effects_df = pd.read_csv(
os.path.join(output_dir, "simulated_muteffects.csv")
)
func_scores = pd.read_csv(
os.path.join(output_dir, "simulated_func_scores.csv")
)
func_scores["func_score_type"] = pd.Categorical(
func_scores["func_score_type"],
categories=["observed_phenotype", "loose_bottle", "tight_bottle"],
ordered=True,
)
# Display-friendly titles for measurement types (used in all subplot titles)
NICE_TITLES = {
"observed_phenotype": "Observed Phenotype",
"loose_bottle": "Loose Bottleneck",
"tight_bottle": "Tight Bottleneck",
}
# Canonical ordering for measurement types
MT_ORDER = ["observed_phenotype", "loose_bottle", "tight_bottle"]
print(
f"Loaded {len(fit_collection_df)} fitted models, "
f"{len(mut_effects_df)} mutations, "
f"{len(func_scores)} functional scores"
)
Loaded 42 fitted models, 1000 mutations, 349866 functional scores
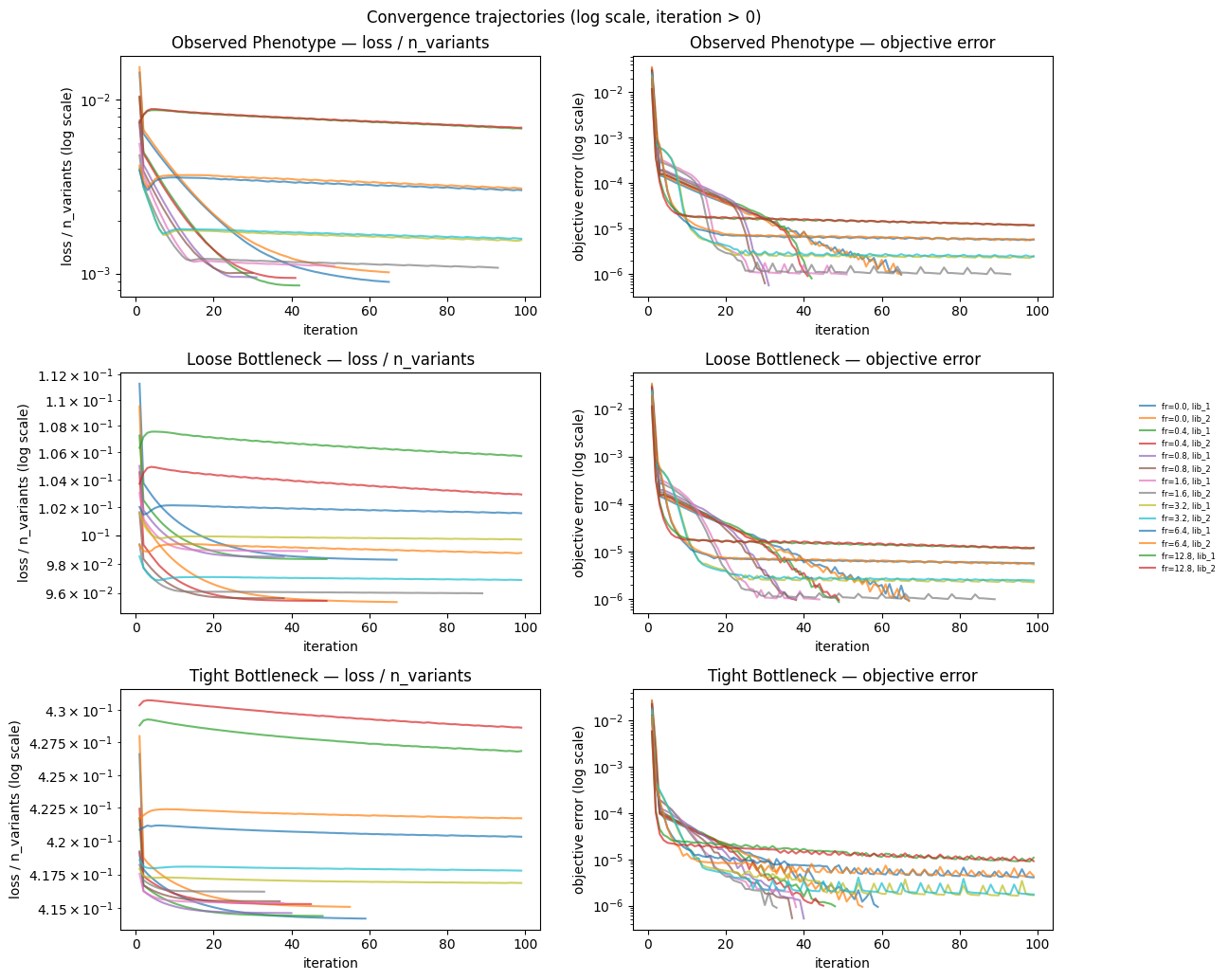
1. Convergence diagnostics¶
[6]:
model_collection = ModelCollection(fit_collection_df)
print("Convergence counts:")
print(model_collection.fit_models.converged.value_counts())
Convergence counts:
converged
True 24
False 18
Name: count, dtype: int64
[7]:
conv_data = model_collection.convergence_trajectory_df(
id_vars=("fusionreg", "measurement_type", "library")
)
conv_data.index.name = "step"
conv_data.reset_index(inplace=True)
# Compute n_variants per model so we can normalize loss to per-variant scale.
# Each (fusionreg, measurement_type, library) combination corresponds to one
# fitted model; we look up its variant count from fit_collection_df.
n_variants_map = {
(row.fusionreg, row.measurement_type, row.library): len(
row.model.data.variants_df
)
for _, row in fit_collection_df.iterrows()
}
conv_data["n_variants"] = conv_data.apply(
lambda r: n_variants_map[
(r["fusionreg"], r["measurement_type"], r["library"])
],
axis=1,
)
# Normalize: total loss divided by number of variants
conv_data["loss_per_variant"] = (
conv_data["loss_trajectory"] / conv_data["n_variants"]
)
# Log-scale convergence trajectory (iteration > 0)
plot_df_log = conv_data.query("iteration > 0")
measurement_types_all = [
mt for mt in MT_ORDER
if mt in plot_df_log["measurement_type"].unique()
]
# Single figure: rows = measurement types, cols = (loss_per_variant, objective_error)
fig, axes = plt.subplots(
len(measurement_types_all), 2,
figsize=(12, 3.5 * len(measurement_types_all)),
squeeze=False,
)
for row_idx, mt in enumerate(measurement_types_all):
sub = plot_df_log[plot_df_log["measurement_type"] == mt]
# Col 0: loss_per_variant
ax0 = axes[row_idx, 0]
for (fr, _, lib), grp in sub.groupby(
["fusionreg", "measurement_type", "library"]
):
ax0.semilogy(
grp["iteration"],
grp["loss_per_variant"],
label=f"fr={fr}, {lib}",
alpha=0.7,
)
ax0.set_title(f"{NICE_TITLES.get(mt, mt)} — loss / n_variants")
ax0.set_xlabel("iteration")
ax0.set_ylabel("loss / n_variants (log scale)")
# Col 1: objective_error_trajectory
ax1 = axes[row_idx, 1]
for (fr, _, lib), grp in sub.groupby(
["fusionreg", "measurement_type", "library"]
):
ax1.semilogy(
grp["iteration"],
grp["objective_error_trajectory"],
label=f"fr={fr}, {lib}",
alpha=0.7,
)
ax1.set_title(f"{NICE_TITLES.get(mt, mt)} — objective error")
ax1.set_xlabel("iteration")
ax1.set_ylabel("objective error (log scale)")
# Remove per-subplot legends and add single consolidated legend
for ax in axes.flat:
leg = ax.get_legend()
if leg:
leg.remove()
handles, labels = axes[0, 0].get_legend_handles_labels()
fig.legend(
handles, labels, loc="center left", bbox_to_anchor=(1.0, 0.5),
frameon=False, fontsize=6,
)
fig.suptitle("Convergence trajectories (log scale, iteration > 0)")
fig.tight_layout(w_pad=3)
fig.subplots_adjust(right=0.93)
plt.show()
2. Mutation effect accuracy (beta and shift vs ground truth)¶
[8]:
model_collection = ModelCollection(fit_collection_df)
groupby = ("library", "measurement_type", "fusionreg")
collection_muts_df = (
model_collection.split_apply_combine_muts(groupby=groupby)
.reset_index()
.rename(
{"beta_h1": "predicted_beta", "shift_h2": "predicted_shift_h2"},
axis=1,
)
.merge(
mut_effects_df.rename(
{
"beta_h1": "true_beta",
"beta_h2": "true_beta_h2",
"shift": "true_shift",
},
axis=1,
),
on="mutation",
)
)
print(f"collection_muts_df: {collection_muts_df.shape[0]} rows")
cache miss - this could take a moment
collection_muts_df: 42000 rows
[9]:
def series_corr(y_true, y_pred):
"""Pearson correlation between arrays."""
return np.corrcoef(y_true, y_pred)[0, 1]
def series_mae(y_true, y_pred):
"""Mean absolute error between arrays."""
return np.mean(np.abs(y_true - y_pred))
new_fit_models_cols = defaultdict(list)
for group, model_mutations_df in collection_muts_df.groupby(list(groupby)):
for i, attribute in enumerate(group):
new_fit_models_cols[groupby[i]].append(group[i])
for parameter in ["beta", "shift"]:
for metric_fxn, name in zip(
[series_corr, series_mae], ["corr", "mae"]
):
postfix = "_h2" if parameter == "shift" else ""
y_pred = model_mutations_df[f"predicted_{parameter}{postfix}"]
y_true = model_mutations_df[f"true_{parameter}"]
new_fit_models_cols[f"{parameter}_{name}"].append(
metric_fxn(y_true, y_pred)
)
model_collection.fit_models = model_collection.fit_models.merge(
pd.DataFrame(new_fit_models_cols), on=list(groupby)
)
[10]:
metric = "corr"
beta_shift_data = (
model_collection.fit_models.assign(
measurement_library=lambda x: (
x["measurement_type"].astype(str) + " " + x["library"]
)
).melt(
id_vars=list(groupby) + ["measurement_library"],
value_vars=[f"beta_{metric}", f"shift_{metric}"],
var_name="parameter",
value_name=metric,
)
)
beta_shift_data["measurement_type"] = pd.Categorical(
beta_shift_data["measurement_type"],
categories=["observed_phenotype", "loose_bottle", "tight_bottle"],
ordered=True,
)
beta_shift_data["parameter"] = (
beta_shift_data["parameter"].str.replace(f"_{metric}", "")
)
beta_shift_data["parameter"] = pd.Categorical(
beta_shift_data["parameter"],
categories=["shift", "beta"],
ordered=True,
)
beta_shift_data["fusionreg_cat"] = beta_shift_data["fusionreg"].astype(str)
for parameter, parameter_df in beta_shift_data.groupby(
"parameter", observed=True
):
mt_vals = [
mt for mt in MT_ORDER
if mt in parameter_df["measurement_type"].dropna().values
]
fig, axes = plt.subplots(
1, len(mt_vals), figsize=(12, 3.5), sharey=True, squeeze=False
)
for ax, mt in zip(axes[0], mt_vals):
sub = parameter_df[parameter_df["measurement_type"] == mt]
for lib, lib_df in sub.groupby("library"):
lib_df = lib_df.sort_values("fusionreg")
marker = "o" if lib == "lib_1" else "s"
ax.plot(
lib_df["fusionreg_cat"],
lib_df[metric],
marker=marker,
label=lib,
)
ax.set_title(NICE_TITLES.get(mt, mt), fontsize=9)
ax.set_xlabel("lasso penalty (\u03bb)")
ax.set_ylabel("Correlation")
ax.tick_params(axis="x", rotation=90)
# Remove per-subplot legends and add single consolidated legend
for ax in axes[0]:
leg = ax.get_legend()
if leg:
leg.remove()
handles, labels = axes[0][0].get_legend_handles_labels()
fig.legend(
handles, labels, loc="center left", bbox_to_anchor=(1.0, 0.5),
frameon=False, fontsize=8,
)
fig.suptitle(
f"Prediction vs. Ground Truth: {parameter}", fontsize=11
)
fig.tight_layout(w_pad=3)
fig.subplots_adjust(right=0.93)
plt.show()
beta_shift_data.to_csv(
os.path.join(output_dir, "model_vs_truth_beta_shift.csv"), index=False
)
print(f"Saved model_vs_truth_beta_shift.csv ({len(beta_shift_data)} rows)")
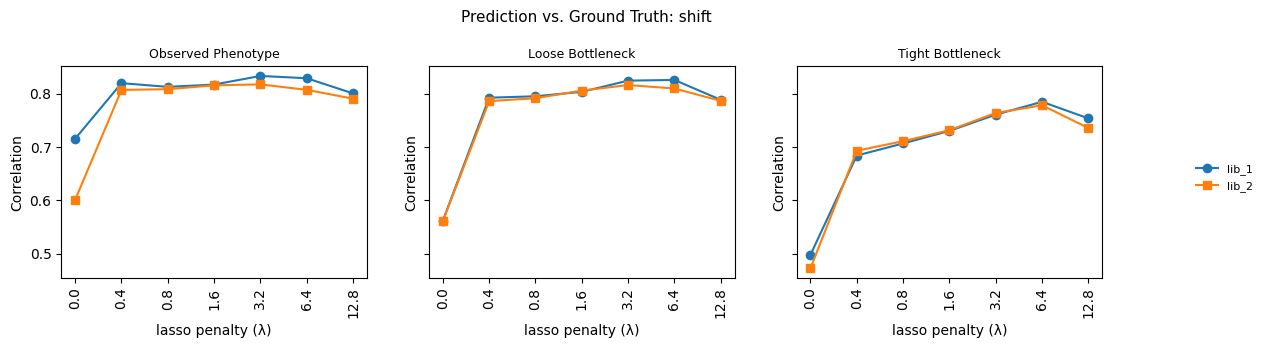
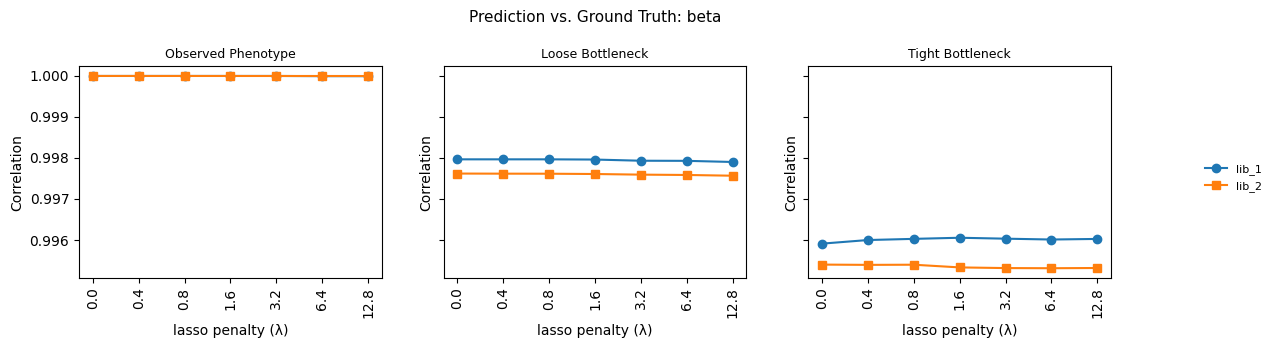
Saved model_vs_truth_beta_shift.csv (84 rows)
[11]:
# Save per-mutation predictions merged with ground truth for downstream figures
collection_muts_df.to_csv(
os.path.join(output_dir, "collection_muts.csv"), index=False
)
print(f"Saved collection_muts.csv ({len(collection_muts_df)} rows)")
Saved collection_muts.csv (42000 rows)
3. Shift sparsity¶
[12]:
_, sparsity_data = model_collection.shift_sparsity(return_data=True)
sparsity_data = sparsity_data.assign(
library=(
sparsity_data.dataset_name.str.split("_").str[:2].str.join("_")
),
library_type=(
sparsity_data.dataset_name.str.split("_").str[:2].str.join("_")
+ "-"
+ sparsity_data.mut_type
),
measurement_type=(
sparsity_data.dataset_name.str.split("_").str[2:4].str.join("_")
),
)
sparsity_data["measurement_type"] = pd.Categorical(
sparsity_data["measurement_type"],
categories=["observed_phenotype", "loose_bottle", "tight_bottle"],
ordered=True,
)
sparsity_data["fusionreg_cat"] = sparsity_data["fusionreg"].astype(str)
true_sparsity = 1 - mut_effects_df.shifted_site.mean()
mt_vals = [
mt for mt in MT_ORDER
if mt in sparsity_data["measurement_type"].dropna().values
]
fig, axes = plt.subplots(1, len(mt_vals), figsize=(12, 3.5), sharey=True, squeeze=False)
for ax, mt in zip(axes[0], mt_vals):
sub = sparsity_data[sparsity_data["measurement_type"] == mt]
ax.axhline(true_sparsity, linestyle="--", color="red", label="true")
for (lt, lib), grp in sub.groupby(["library_type", "library"]):
grp = grp.sort_values("fusionreg")
marker = "o" if lib == "lib_1" else "s"
ax.plot(
grp["fusionreg_cat"],
grp["sparsity"],
marker=marker,
label=lt,
alpha=0.8,
)
ax.set_title(NICE_TITLES.get(mt, mt), fontsize=9)
ax.set_xlabel("lasso penalty (\u03bb)")
ax.set_ylabel("sparsity")
ax.tick_params(axis="x", rotation=90)
# Remove per-subplot legends and add single consolidated legend
for ax in axes[0]:
leg = ax.get_legend()
if leg:
leg.remove()
handles, labels = axes[0][0].get_legend_handles_labels()
fig.legend(
handles, labels, loc="center left", bbox_to_anchor=(1.0, 0.5),
frameon=False, fontsize=7,
)
fig.suptitle("Shift sparsity vs ground truth", fontsize=11)
fig.tight_layout(w_pad=3)
fig.subplots_adjust(right=0.93)
plt.show()
sparsity_data.to_csv(
os.path.join(output_dir, "fit_sparsity.csv"), index=False
)
print(f"Saved fit_sparsity.csv ({len(sparsity_data)} rows)")
cache miss - this could take a moment
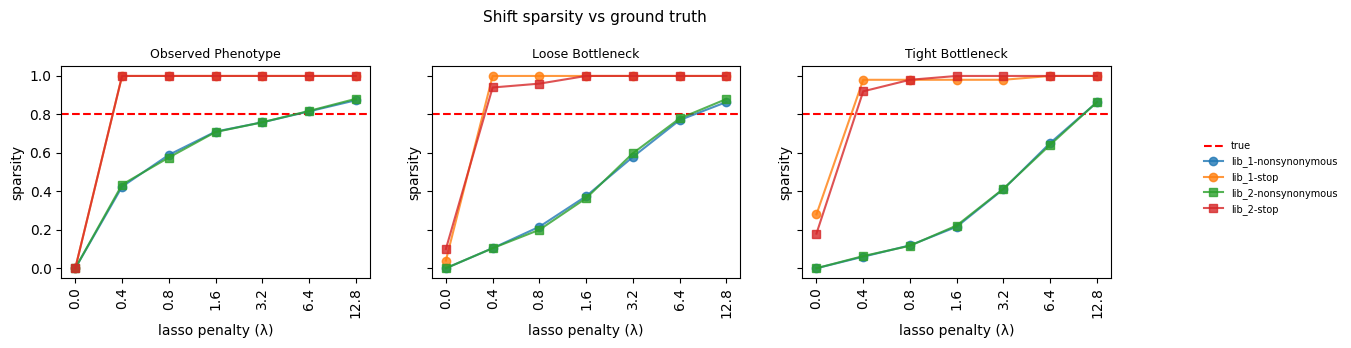
Saved fit_sparsity.csv (84 rows)
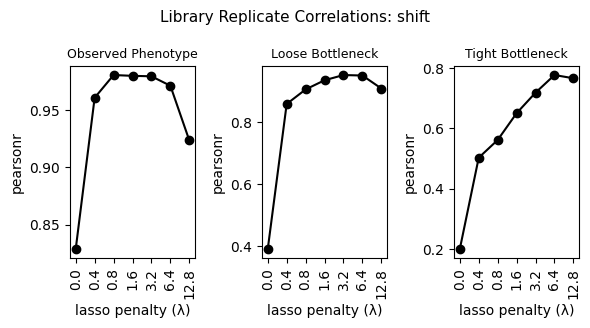
4. Replicate correlations¶
[13]:
_, corr_data = model_collection.mut_param_dataset_correlation(
return_data=True
)
corr_data = (
corr_data.assign(
lib1=corr_data.datasets.str.split(",").str[0],
lib2=corr_data.datasets.str.split(",").str[1],
measurement_type_1=lambda x: (
x["lib1"].str.split("_").str[2:4].str.join("_")
),
measurement_type_2=lambda x: (
x["lib2"].str.split("_").str[2:4].str.join("_")
),
)
.query(
"(measurement_type_1 == measurement_type_2) "
"& (~mut_param.str.contains('predicted_func_score'))"
)
.rename({"measurement_type_1": "measurement_type"}, axis=1)
.replace({"shift_h2": "shift"})
.drop(["lib1", "lib2", "datasets", "measurement_type_2"], axis=1)
)
corr_data["measurement_type"] = pd.Categorical(
corr_data["measurement_type"],
categories=["observed_phenotype", "loose_bottle", "tight_bottle"],
ordered=True,
)
corr_data["mut_param"] = pd.Categorical(
corr_data["mut_param"],
categories=["shift", "beta"],
ordered=True,
)
corr_data["fusionreg_cat"] = corr_data["fusionreg"].astype(str)
for parameter, parameter_df in corr_data.groupby(
"mut_param", observed=True
):
mt_vals = [
mt for mt in MT_ORDER
if mt in parameter_df["measurement_type"].dropna().values
]
fig, axes = plt.subplots(
1, len(mt_vals), figsize=(6, 3.3), squeeze=False
)
for ax, mt in zip(axes[0], mt_vals):
sub = parameter_df[
parameter_df["measurement_type"] == mt
].sort_values("fusionreg")
ax.plot(
sub["fusionreg_cat"], sub["correlation"], marker="o", color="k"
)
ax.set_title(NICE_TITLES.get(mt, mt), fontsize=9)
ax.set_xlabel("lasso penalty (\u03bb)")
ax.set_ylabel("pearsonr")
ax.tick_params(axis="x", rotation=90)
fig.suptitle(
f"Library Replicate Correlations: {parameter}", fontsize=11
)
fig.tight_layout()
plt.show()
corr_data.to_csv(
os.path.join(output_dir, "library_replicate_correlation.csv"),
index=False,
)
print(
f"Saved library_replicate_correlation.csv ({len(corr_data)} rows)"
)
Saved library_replicate_correlation.csv (84 rows)
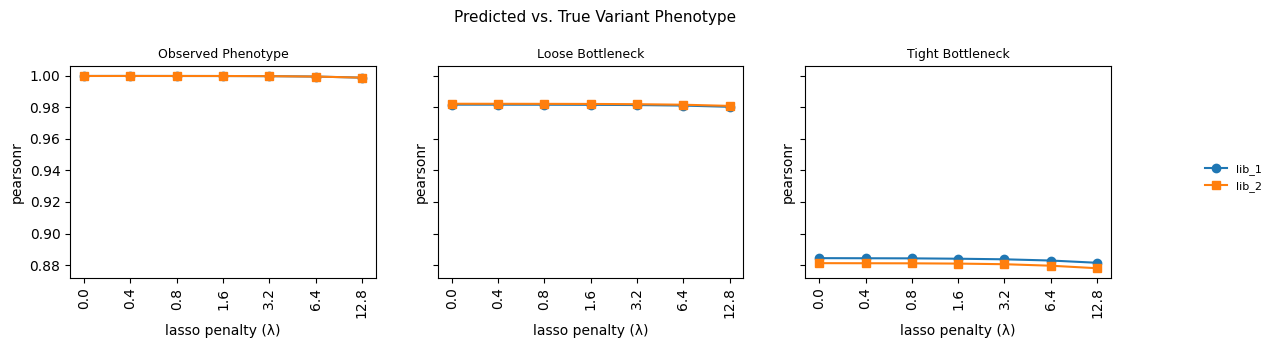
5. Variant phenotype predictions¶
[14]:
phenotype_fxn_dict_h1, phenotype_fxn_dict_h2 = reconstruct_simulators(
sim, mut_effects_df, seed
)
print("Phenotype functions reconstructed")
Phenotype functions reconstructed
[15]:
variants_df = pd.concat(
[
row.model.get_variants_df(phenotype_as_effect=False)
.assign(
library=row.library,
measurement_type=row.measurement_type,
fusionreg=row.fusionreg,
)
.rename(
{
"predicted_func_score": "predicted_phenotype",
"predicted_latent": "predicted_latent_phenotype",
"func_score": "measured_phenotype",
},
axis=1,
)
.assign(
predicted_enrichment=lambda x: 2 ** x["predicted_phenotype"],
measured_enrichment=lambda x: 2 ** x["measured_phenotype"],
fit_idx=idx,
)
for idx, row in fit_collection_df.iterrows()
]
)
# Add ground truth phenotypes from reconstructed simulators
variants_df = pd.concat(
[
variants_df.query("condition == @homolog").assign(
true_latent_phenotype=lambda x: x["aa_substitutions"].apply(
pfxn["latentPhenotype"]
),
true_observed_phenotype=lambda x: x["aa_substitutions"].apply(
pfxn["observedPhenotype"]
),
true_enrichment=lambda x: x["aa_substitutions"].apply(
pfxn["observedEnrichment"]
),
)
for homolog, pfxn in [
("h1", phenotype_fxn_dict_h1),
("h2", phenotype_fxn_dict_h2),
]
]
)
variants_df["measurement_type"] = pd.Categorical(
variants_df["measurement_type"],
categories=["observed_phenotype", "loose_bottle", "tight_bottle"],
ordered=True,
)
print(f"Variant predictions: {len(variants_df)} rows")
Variant predictions: 2449062 rows
[16]:
# Add variant phenotype metrics to fit_collection_df
for idx, model_variants_df in variants_df.groupby("fit_idx"):
for metric_fxn, metric_name in zip(
[series_corr, series_mae], ["corr", "mae"]
):
fit_collection_df.loc[
idx, f"variant_phenotype_{metric_name}"
] = metric_fxn(
model_variants_df["measured_phenotype"],
model_variants_df["predicted_phenotype"],
)
pheno_data = fit_collection_df[
[
"library",
"measurement_type",
"fusionreg",
"dataset_name",
"variant_phenotype_corr",
"variant_phenotype_mae",
]
].copy()
pheno_data["fusionreg_cat"] = pheno_data["fusionreg"].astype(str)
# Plot variant phenotype correlation vs fusionreg
mt_vals = [
mt for mt in MT_ORDER
if mt in pheno_data["measurement_type"].dropna().values
]
fig, axes = plt.subplots(1, len(mt_vals), figsize=(12, 3.5), sharey=True, squeeze=False)
for ax, mt in zip(axes[0], mt_vals):
sub = pheno_data[pheno_data["measurement_type"] == mt]
for lib, lib_df in sub.groupby("library"):
lib_df = lib_df.sort_values("fusionreg")
marker = "o" if lib == "lib_1" else "s"
ax.plot(
lib_df["fusionreg_cat"],
lib_df["variant_phenotype_corr"],
marker=marker,
label=lib,
)
ax.set_title(NICE_TITLES.get(mt, mt), fontsize=9)
ax.set_xlabel("lasso penalty (\u03bb)")
ax.set_ylabel("pearsonr")
ax.tick_params(axis="x", rotation=90)
# Remove per-subplot legends and add single consolidated legend
for ax in axes[0]:
leg = ax.get_legend()
if leg:
leg.remove()
handles, labels = axes[0][0].get_legend_handles_labels()
fig.legend(
handles, labels, loc="center left", bbox_to_anchor=(1.0, 0.5),
frameon=False, fontsize=8,
)
fig.suptitle("Predicted vs. True Variant Phenotype", fontsize=11)
fig.tight_layout(w_pad=3)
fig.subplots_adjust(right=0.93)
plt.show()
pheno_data.to_csv(
os.path.join(output_dir, "model_vs_truth_variant_phenotype.csv"),
index=False,
)
print(
f"Saved model_vs_truth_variant_phenotype.csv ({len(pheno_data)} rows)"
)
Saved model_vs_truth_variant_phenotype.csv (42 rows)
7. Additional visualizations¶
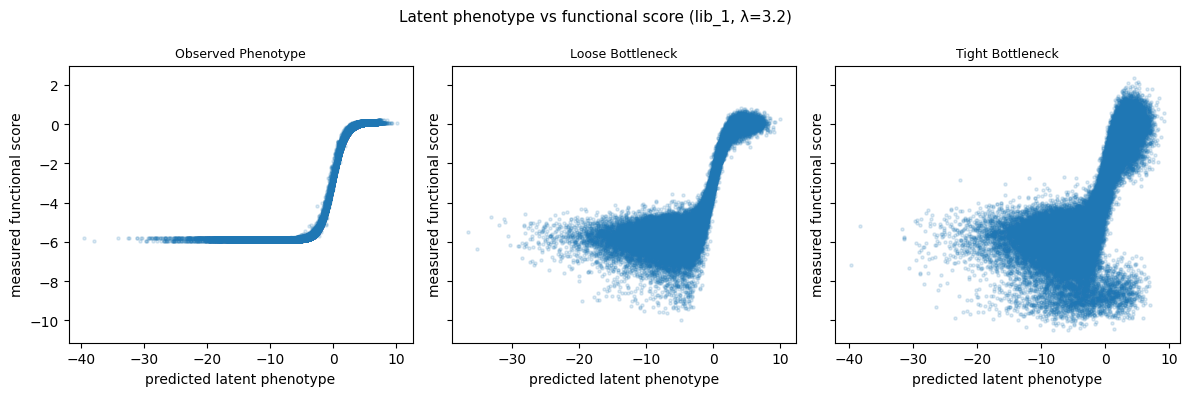
Latent phenotype vs functional score¶
Scatter of predicted latent phenotype vs measured functional score, faceted by measurement type. Uses the chosen lasso value and lib_1.
[17]:
plot_variants = variants_df.query(
f"library == 'lib_1' & fusionreg == {lasso_choice}"
)
mt_vals = [
mt for mt in MT_ORDER
if mt in plot_variants["measurement_type"].dropna().values
]
fig, axes = plt.subplots(1, len(mt_vals), figsize=(12, 4), sharey=True, squeeze=False)
for ax, mt in zip(axes[0], mt_vals):
sub = plot_variants[plot_variants["measurement_type"] == mt]
ax.scatter(
sub["predicted_latent_phenotype"],
sub["measured_phenotype"],
alpha=0.15,
s=5,
rasterized=True,
)
ax.set_xlabel("predicted latent phenotype")
ax.set_ylabel("measured functional score")
ax.set_title(NICE_TITLES.get(mt, mt), fontsize=9)
fig.suptitle(
f"Latent phenotype vs functional score (lib_1, \u03bb={lasso_choice})",
fontsize=11,
)
fig.tight_layout()
plt.show()
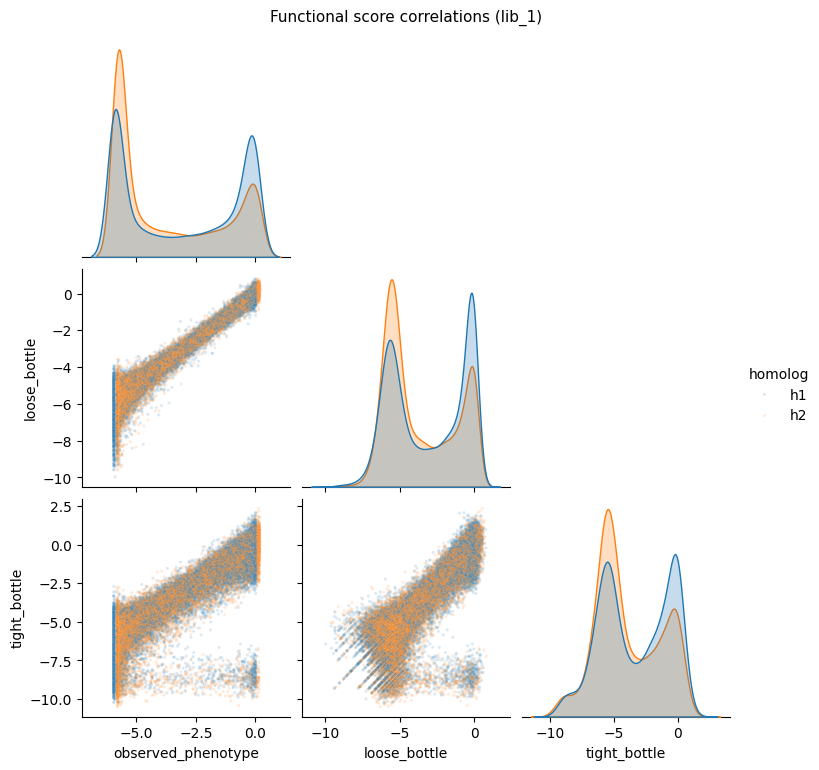
Functional score pairplot¶
Seaborn pairplot showing correlations of functional scores across measurement types for lib_1.
[18]:
# Pivot functional scores for pairplot
pairplot_df = (
func_scores.query("library == 'lib_1'")
.pivot_table(
index=["aa_substitutions", "homolog"],
columns="func_score_type",
values="func_score",
)
.dropna()
.reset_index()
)
score_cols = [
c for c in pairplot_df.columns
if c not in ["aa_substitutions", "homolog"]
]
g = sns.pairplot(
pairplot_df,
vars=score_cols,
hue="homolog",
plot_kws={"alpha": 0.15, "s": 5, "rasterized": True},
diag_kind="kde",
height=2.5,
corner=True,
)
g.figure.suptitle(
"Functional score correlations (lib_1)", y=1.02, fontsize=11
)
# Move legend outside and remove frame
if g._legend is not None:
g._legend.set_bbox_to_anchor((1.0, 0.5))
g._legend.set_frame_on(False)
plt.show()
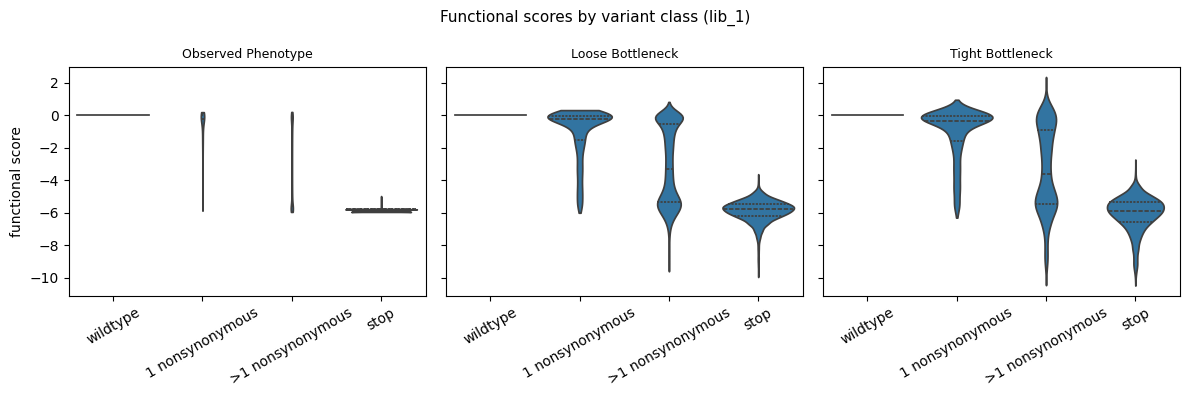
Functional score distribution by variant class¶
Violin plot of functional scores stratified by the number of mutations per variant.
[19]:
violin_df = func_scores.query("library == 'lib_1'").copy()
# Classify variants by mutation count
def classify_variant(aa_subs):
"""Classify variant by mutation type."""
if pd.isna(aa_subs) or aa_subs.strip() == "":
return "wildtype"
elif "*" in aa_subs:
return "stop"
elif len(aa_subs.split()) == 1:
return "1 nonsynonymous"
else:
return ">1 nonsynonymous"
violin_df["variant_class"] = violin_df["aa_substitutions"].apply(
classify_variant
)
class_order = ["wildtype", "1 nonsynonymous", ">1 nonsynonymous", "stop"]
violin_df["variant_class"] = pd.Categorical(
violin_df["variant_class"], categories=class_order, ordered=True
)
mt_vals = [
mt for mt in MT_ORDER
if mt in violin_df["func_score_type"].dropna().values
]
fig, axes = plt.subplots(1, len(mt_vals), figsize=(12, 4), sharey=True, squeeze=False)
for ax, mt in zip(axes[0], mt_vals):
sub = violin_df[violin_df["func_score_type"] == mt]
sns.violinplot(
data=sub,
x="variant_class",
y="func_score",
ax=ax,
cut=0,
inner="quartile",
)
ax.set_title(NICE_TITLES.get(mt, mt), fontsize=9)
ax.set_xlabel("")
ax.set_ylabel("functional score")
ax.tick_params(axis="x", rotation=30)
fig.suptitle(
"Functional scores by variant class (lib_1)", fontsize=11
)
fig.tight_layout()
plt.show()
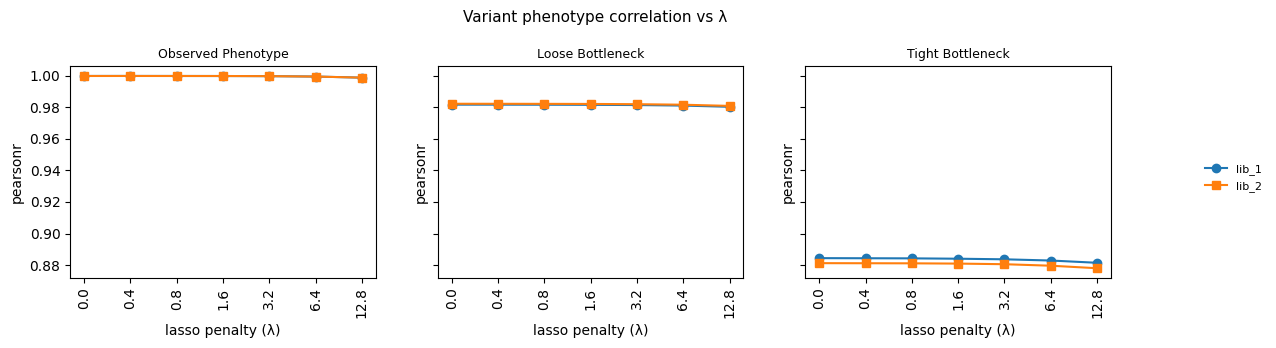
Variant phenotype correlation vs lambda¶
Line plot of variant phenotype correlation across the fusionreg grid, faceted by measurement type.
[20]:
# Use the pheno_data already computed in section 5
mt_vals = [
mt for mt in MT_ORDER
if mt in pheno_data["measurement_type"].dropna().values
]
fig, axes = plt.subplots(
1, len(mt_vals), figsize=(12, 3.5), sharey=True, squeeze=False
)
for ax, mt in zip(axes[0], mt_vals):
sub = pheno_data[pheno_data["measurement_type"] == mt]
for lib, lib_df in sub.groupby("library"):
lib_df = lib_df.sort_values("fusionreg")
marker = "o" if lib == "lib_1" else "s"
ax.plot(
lib_df["fusionreg_cat"],
lib_df["variant_phenotype_corr"],
marker=marker,
label=lib,
)
ax.set_title(NICE_TITLES.get(mt, mt), fontsize=9)
ax.set_xlabel("lasso penalty (\u03bb)")
ax.set_ylabel("pearsonr")
ax.tick_params(axis="x", rotation=90)
# Remove per-subplot legends and add single consolidated legend
for ax in axes[0]:
leg = ax.get_legend()
if leg:
leg.remove()
handles, labels = axes[0][0].get_legend_handles_labels()
fig.legend(
handles, labels, loc="center left", bbox_to_anchor=(1.0, 0.5),
frameon=False, fontsize=8,
)
fig.suptitle(
"Variant phenotype correlation vs \u03bb", fontsize=11
)
fig.tight_layout(w_pad=3)
fig.subplots_adjust(right=0.93)
plt.show()
Diagnostic Visualizations¶
Inline diagnostic plots adapted from the simulation pipeline. No files are saved.
Fitted model GE landscape¶
Use the model’s built-in GE landscape plot for one representative fit (chosen lasso, loose_bottle, lib_1).
[21]:
representative_fit = (
model_collection.fit_models
.query(
f"fusionreg == {lasso_choice} "
"and measurement_type == 'loose_bottle' "
"and library == 'lib_1'"
)
.iloc[0]
)
multidms.plot.ge_landscape(representative_fit.model)
[21]: