Cross-Validation¶
Evaluate model generalization via 80/20 train/test cross-validation across the fusion-regularization grid.
Outline
Load training functional scores and pipeline config
Split data into train (80%) and test (20%) sets per replicate
Fit models on training data across the fusionreg grid
Evaluate validation loss on held-out test data
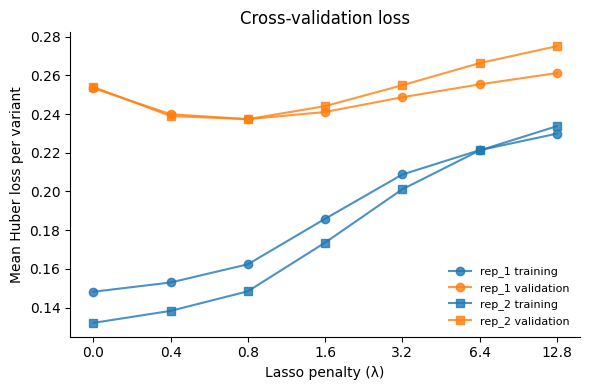
Visualize training vs validation loss
[1]:
import warnings
warnings.filterwarnings("ignore")
import os
import sys
sys.path.insert(0, "notebooks")
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import multidms
from multidms.model_collection import ModelCollection, fit_models
from multidms.utils import explode_params_dict
from _common import load_config, build_fit_params
[2]:
config_path = "config/config.yaml"
[3]:
# Parameters
config_path = "config/config.yaml"
[4]:
config = load_config(config_path)
spike = config["spike"]
fit_config = spike["fitting"]
reference = spike["reference"]
seed = config["seed"]
train_frac = config["train_frac"]
condition_titles = spike["condition_titles"]
output_dir = "results"
os.makedirs(output_dir, exist_ok=True)
Load training functional scores¶
[5]:
func_score_df = pd.read_csv(
os.path.join(output_dir, "training_functional_scores.csv")
).fillna({"aa_substitutions": ""})
print(f"Loaded {len(func_score_df):,} variants")
Loaded 281,158 variants
Train/test split¶
Split functional scores 80/20 per replicate. Wildtype variants are kept in the training set.
[6]:
train_datasets = []
test_data = {}
for rep_num, df_rep in func_score_df.groupby("replicate"):
# Aggregate per (condition, aa_substitutions)
df_agg = (
df_rep.groupby(["condition", "aa_substitutions"], dropna=False)
.agg({"func_score": "mean"})
.reset_index()
)
wt_mask = df_agg["aa_substitutions"].str.strip() == ""
wt_rows = df_agg[wt_mask]
non_wt_rows = df_agg[~wt_mask].sample(frac=1, random_state=seed)
n_train = int(len(non_wt_rows) * train_frac)
train_split = pd.concat([wt_rows, non_wt_rows.iloc[:n_train]])
test_split = non_wt_rows.iloc[n_train:]
name = f"rep_{rep_num}"
train_datasets.append(
multidms.Data(
train_split,
reference=reference,
alphabet=multidms.AAS_WITHSTOP_WITHGAP,
assert_site_integrity=False,
verbose=False,
name=name,
)
)
test_data[name] = test_split
print(f"{name}: {len(train_split):,} train, {len(test_split):,} test")
rep_1: 116,776 train, 29,194 test
rep_2: 108,151 train, 27,037 test
Fit CV models¶
[7]:
cv_fitting_params = build_fit_params(fit_config, train_datasets)
n_models = len(explode_params_dict(cv_fitting_params))
cfg_n_processes = fit_config.get("n_processes")
if cfg_n_processes is None:
n_processes = min(os.cpu_count() // 2, n_models)
else:
n_processes = min(int(cfg_n_processes), n_models)
n_processes = max(n_processes, 1)
print(f"Fitting {n_models} CV models with n_processes={n_processes}")
n_fit, n_failed, fit_collection_cv = fit_models(
cv_fitting_params, n_processes=n_processes
)
for col in fit_collection_cv.columns:
if fit_collection_cv[col].apply(lambda x: isinstance(x, dict)).any():
fit_collection_cv[col] = fit_collection_cv[col].apply(str)
print(f"CV: fit {n_fit} models, {n_failed} failed")
Fitting 14 CV models with n_processes=14
CV: fit 14 models, 0 failed
Evaluate validation loss¶
[8]:
mc_cv = ModelCollection(fit_collection_cv)
# Filter test variants to those with known mutations only
filtered_test = {}
for name, test_df in test_data.items():
model = (
mc_cv.fit_models.query(f"dataset_name == '{name}'")
.iloc[0]
.model
)
known_muts = set(model.data.mutations)
def all_muts_known(subs):
if pd.isna(subs) or subs.strip() == "":
return True
return all(m in known_muts for m in subs.split())
mask = test_df["aa_substitutions"].apply(all_muts_known)
filtered_test[name] = test_df[mask]
n_dropped = (~mask).sum()
if n_dropped > 0:
print(f"{name}: dropped {n_dropped}/{len(test_df)} test variants with unseen mutations")
mc_cv.add_eval_loss(filtered_test, overwrite=True)
rep_1: dropped 5447/29194 test variants with unseen mutations
rep_2: dropped 4999/27037 test variants with unseen mutations
Visualize CV loss¶
[9]:
cv_data = mc_cv.fit_models.copy()
cv_data = (
cv_data.melt(
id_vars=["fusionreg", "dataset_name"],
value_vars=["total_loss_training", "total_loss_validation"],
var_name="dataset",
value_name="loss",
)
.assign(dataset=lambda x: x["dataset"].str.replace("total_loss_", ""))
)
# Normalize loss by sample count
n_samples = {}
for d in train_datasets:
n_train = sum(
len(d.variants_df.query(f"condition == '{c}'"))
for c in d.conditions
)
n_samples[(d.name, "training")] = n_train
for name, test_df in filtered_test.items():
n_test = sum(
len(test_df.query(f"condition == '{c}'"))
for c in test_df["condition"].unique()
)
n_samples[(name, "validation")] = n_test
cv_data["n_samples"] = cv_data.apply(
lambda row: n_samples.get((row["dataset_name"], row["dataset"]), 1),
axis=1,
)
cv_data["mean_loss"] = cv_data["loss"] / cv_data["n_samples"]
cv_data["fusionreg_cat"] = cv_data["fusionreg"].astype(str)
# Plot
fig, ax = plt.subplots(figsize=(6, 4))
for (rep, ds), grp in cv_data.groupby(["dataset_name", "dataset"]):
grp = grp.sort_values("fusionreg")
marker = "o" if "rep_1" in rep else "s"
color = "tab:blue" if ds == "training" else "tab:orange"
ax.plot(
grp["fusionreg_cat"], grp["mean_loss"],
marker=marker, color=color, alpha=0.8,
label=f"{rep} {ds}",
)
ax.set_xlabel("Lasso penalty (\u03bb)")
ax.set_ylabel("Mean Huber loss per variant")
ax.set_title("Cross-validation loss")
ax.legend(fontsize=8, frameon=False)
ax.spines[["top", "right"]].set_visible(False)
plt.tight_layout()
plt.show()
Save¶
[10]:
cv_data.to_csv(
os.path.join(output_dir, "cross_validation_loss.csv"), index=False
)
print(f"Saved cross_validation_loss.csv ({len(cv_data)} rows)")
Saved cross_validation_loss.csv (28 rows)
